Revolutionizing Gene Therapy: Screening Lipid Nanoparticles for optimal delivery of mRNA and CRISPR-Cas9
The world of medical science is witnessing groundbreaking and fast paced transformation as researchers delve into the potential of lipid nanoparticles (LNPs) for the delivery of messenger RNA (mRNA) and gene editing material.
Researchers at MIT and the University of Toronto have recently released a thorough study at the intersection of LNPs, mRNA, and genome editing, offering insights into how these innovative nanoparticles are poised to reshape the landscape of precision medicine.
The power of LNPs for mRNA Delivery and gene editing
Lipid nanoparticles have emerged as game-changing carriers for delivering mRNA and CRISPR-Cas9 directly into cells.
Thanks to the integration of an ionizable lipid into their composition, LNPs offer the unique capabilities of changing structure depending on the pH, and thus location in the body they are at, to deliver their encapsulated material to the desired location. Check out our review on Lipid Nanoparticle to learnmore!
This capability is particularly promising in the realm of gene therapy, where the ability to introduce specific genetic material directly into cells can lead to targeted therapeutic outcomes. By encapsulating mRNA or gene edition material (CRISPR-Cas9) within these biodegradable LNPs, researchers are unlocking the potential to treat a wide array of genetic disorders and diseases.
In this article, they demonstrate the power of that innovative toolbox through an example on Congenital lung disease. Congenital lung diseases have long posed significant health challenges, often linked to genetic factors. While the genetic origins of these diseases are well-known, finding effective treatments has remained elusive.
Traditional gene therapy methods, often utilizing viral vectors, carry the risk of unwanted genetic alterations and immune responses, while offering poor transfection efficiency. Here is where lipid nanoparticles step in. These biodegradable, ionizable LNPs are emerging as a groundbreaking solution to deliver gene-editing tools directly to lung epithelial cells, however they have so far shown poor transfection to airway epithelial cells. The goal of their study is hence to study LNP composition, to enhance and optimize CRISPR-Cas9 delivery to those specific cells.
Revolutionizing LNP Delivery with high volume screening
To harness the potential of LNPs, researchers have developed an innovative approach, combining high-throughput synthesis and screening.
They first started with the synthesis of a broad range of ionizable lipids by combining 72 different amine headgroups with 10 different lipid tails, to create a library of 720 different ionizable lipids.
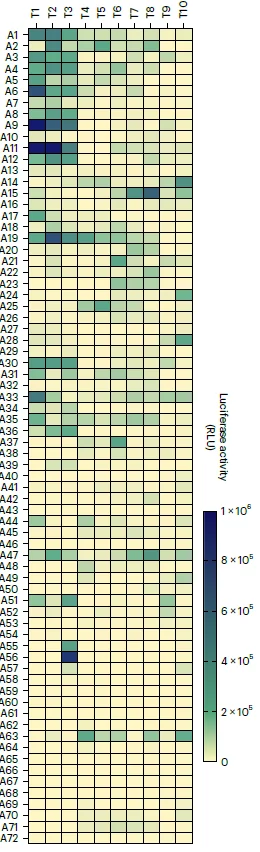
Using this broad set of lipids, they synthesized 720 mRNA-loaded LNP (each comprising DOTAP, PEG, Cholesterol and one of the headgroup/lipid tail combination) encapsulating firefly luciferase (mLuc), which can easily be detected by bioluminescence, with the goal of identifying the most promising ionizable group for delivery of encapsulated payload.
They characterized this set of mLuc-LNP on A549 cells to determine their hit rate. This first round of characterization showed astonishing results as the percentage of hit rate could vary from a nearly null hit rate to very high simply depending on the type of ionizable lipids used, showing once again the importance of testing numerous LNP combinations for the proper delivery of RNA within cells.
These first steps allowed them to identify the 15 most promising ionizable headgroups (those showing average transcription efficiency >80%) for further in-vivo testing.
To minimize animal use and time during in-vivo trials, they carried out orthogonal batch-based testing. To do so, they first tested on mice each of the identified ionizable headgroups with all the possible tails combinations at once, to identify the headgroups offering the best response in-vivo at the injection site, allowing them single out 7 different lipids headgroup candidates. Using those, they tested all the possibles headgroups-tail combination to eventually select the 9 LNP with the highest potential.
Once the LNP candidates were selected by mean of mRNA (mLuc), they investigated their potential to deliver CRISPR-Cas9 complexes on the HEK 293T cell line: It appeared that 6 of the 9 candidates outperformed positive control, with one candidate – the one displaying the highest mLUC transfection – showing the highest efficiency.
Comparison with other FDA-approved LNP: better transcription efficiency and lower toxicity
Once the most promising LNP group for CRISPR-Cas9 was identified, researchers in the study decided to switch to a microfluidics-based synthesis approach, to improve the control of the size of the synthesized nanoparticle, as well as decrease polydispersity and improved encapsulation efficiency. Learn more on the topic in our microfluidic synthesis of nanoparticle review.
Using microfluidic formulation, they compared the mLuc response in lung transfection efficiency of their identified LNP (which they slightly modified to incorporate DOTAP instead of DOPE, as it shows better transcription efficiency) with DLin-MC3-DMA (MC3), an ionizable lipid approved by the US FDA for RNA delivery.
It appeared that their identified LNP combination showed an improved efficiency of approximately 100 folds compared to the FDA-approved one.
Additionally, toxicity was shown to be lowered using their identified LNP as <30% of it would remain in the lung 48 hours after injection, compared with >90% with the FDA-approved LNP.
mRNA delivery by LNP in lungs
To test the efficiency in lungs, the researchers used Cre-recombinase mRNA (Cre-mRNA) to edit genes in mice, so that they produce tdTomato, a red fluorescent protein, and found that the LNPs could efficiently deliver this mRNA, resulting in nearly 50% of the target lung cells expressing fluorescence.
They also noticed that delivery was twice higher than by polymer polyplex, and that multiple doses of LNPs increased the efficiency of gene editing (by 20% for 3 doses instead of 1). The researchers also demonstrated that this approach could target specific cell types in the respiratory epithelium.
CRISPR-Cas9 delivery
The researchers then extended their approach to CRISPR-Cas9 gene editing. They coencapsulated sgRNA (guide RNA) and SpCas9 mRNA (enzymes for gene editing) into the RCB-4-8 LNPs and administered them to mice. They achieved successful gene editing with tdTomato reporter activation, although the efficiency was lower compared to Cre recombinase-mediated editing in the previous experiment. They also explored a combined approach using both LNPs and AAV (adeno-associated virus) for gene editing, showing promising results in terms of efficiency and avoiding potential safety concerns.
Conclusion
Through this study researchers have aimed at optimizing the delivery of mRNA and CRISPR-Cas9 components by synthesized and tested on cells a set of 720 ionizable lipid, to identify one promising candidate which showed transcription efficiency at least 100 times higher than FDA-approved LNP, while offering much lower toxicity.
Using this identified LNP, they tested in-vivo the deliverability of mRNA and gene editing tool in the lung of mouse, and proved that not only the delivery was much improved compared to polymer polyplex, but also that they could also target specific cell types in the respiratory epithelium.
If you want to learn more and go deeper onto the topic, we suggest you to have a look at the full article: https://doi.org/10.1038/s41587-023-01679-x
